The StructProp Class¶
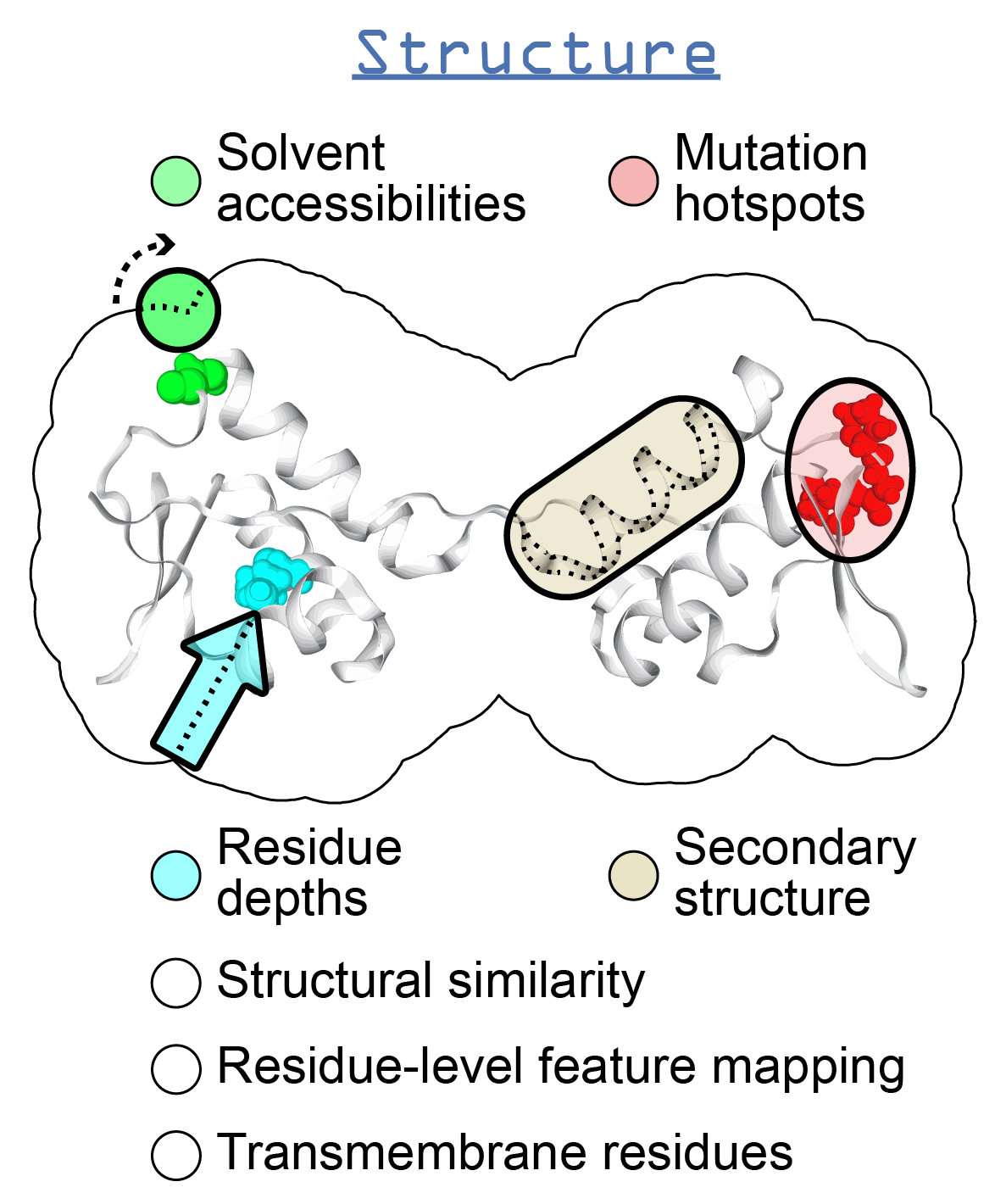
Introduction¶
This section will give an overview of the methods that can be executed for a single protein structure.
Available functions¶
Sequence & structure-based predictions¶
| Function | Description | Internal Python class used and functions provided |
External software to install |
Web server | Alternate external software to install |
|---|---|---|---|---|---|
| Homology modeling | Preparation scripts and parsers for executing homology modeling algorithms |
I-TASSER | |||
| Transmembrane orientation |
Prediction of transmembrane domains and orientation in a membrane |
opm module |
OPM | ||
| Kinetic folding rate | Prediction of protein folding rates from amino acid sequence |
kinetic_folding_rate module |
FOLD-RATE |
Structure-based calculations or functions¶
| Function | Description | Internal Python class used and functions provided |
External software to install |
Web server | Alternate external software to install |
|---|---|---|---|---|---|
| Secondary structure | Calculations of secondary structure | DSSP | STRIDE | ||
| Solvent accessibilities | Calculations of per-residue absolute and relative solvent accessibilities |
DSSP | FreeSASA | ||
| Residue depths | Calculations of residue depths | MSMS | |||
| Structural similarity | Pairwise calculations of 3D structural similarity |
fatcat module |
FATCAT | ||
| Various structure properties |
Basic properties of the structure, such as distance measurements between residues or number of disulfide bridges |
||||
| Quality | Custom functions to allow ranking of structures by percent identity to a defined sequence, structure resolution, and other structure quality metrics |
set_representative_structure function |
|||
| Structure cleaning, mutating |
Custom functions to allow for the preparation of structure files for molecular modeling, with options to remove hydrogens/waters/heteroatoms, select specific chains, or mutate specific residues. |
AmberTools |
API¶
StructProp¶
-
class
ssbio.protein.structure.structprop.StructProp(ident, description=None, chains=None, mapped_chains=None, is_experimental=False, structure_path=None, file_type=None)[source]¶ Generic class to represent information for a protein structure.
The main utilities of this class are to:
- Provide access to the 3D coordinates using a Biopython Structure object through the method
parse_structure. - Run predictions and computations on the structure
- Analyze specific chains using the
mapped_chainsattribute - Provide wrapper methods to
nglviewto view the structure in a Jupyter notebook
Parameters: - ident (str) – Unique identifier for this structure
- description (str) – Optional human-readable description
- chains (str, list) – Chain ID or list of IDs
- mapped_chains (str, list) – A chain ID or IDs to indicate what chains should be analyzed
- is_experimental (bool) – Flag to indicate if structure is an experimental or computational model
- structure_path (str) – Path to structure file
- file_type (str) – Type of structure file -
pdb,pdb.gz,mmcif,cif,cif.gz,xml.gz,mmtf,mmtf.gz
-
add_chain_ids(chains)[source]¶ Add chains by ID into the chains attribute
Parameters: chains (str, list) – Chain ID or list of IDs
-
add_mapped_chain_ids(mapped_chains)[source]¶ Add chains by ID into the mapped_chains attribute
Parameters: mapped_chains (str, list) – Chain ID or list of IDs
-
add_residues_highlight_to_nglview(view, structure_resnums, chain=None, res_color='red')[source]¶ Add a residue number or numbers to an NGLWidget view object.
Parameters: - view (NGLWidget) – NGLWidget view object
- structure_resnums (int, list) – Residue number(s) to highlight, structure numbering
- chain (str, list) – Chain ID or IDs of which residues are a part of. If not provided, all chains in the mapped_chains attribute will be used. If that is also empty, and exception is raised.
- res_color (str) – Color to highlight residues with
-
add_scaled_residues_highlight_to_nglview(view, structure_resnums, chain=None, color='red', unique_colors=False, opacity_range=(0.5, 1), scale_range=(0.7, 10))[source]¶ - Add a list of residue numbers (which may contain repeating residues) to a view, or add a dictionary of
- residue numbers to counts. Size and opacity of added residues are scaled by counts.
Parameters: - view (NGLWidget) – NGLWidget view object
- structure_resnums (int, list, dict) – Residue number(s) to highlight, or a dictionary of residue number to frequency count
- chain (str, list) – Chain ID or IDs of which residues are a part of. If not provided, all chains in the mapped_chains attribute will be used. If that is also empty, and exception is raised.
- color (str) – Color to highlight residues with
- unique_colors (bool) – If each mutation should be colored uniquely (will override color argument)
- opacity_range (tuple) – Min/max opacity values (residues that have higher frequency counts will be opaque)
- scale_range (tuple) – Min/max size values (residues that have higher frequency counts will be bigger)
-
chains= None¶ DictList – A DictList of chains have their sequence stored in them, along with residue-specific
-
clean_structure(out_suffix='_clean', outdir=None, force_rerun=False, remove_atom_alt=True, keep_atom_alt_id='A', remove_atom_hydrogen=True, add_atom_occ=True, remove_res_hetero=True, keep_chemicals=None, keep_res_only=None, add_chain_id_if_empty='X', keep_chains=None)[source]¶ Clean the structure file associated with this structure, and save it as a new file. Returns the file path.
Parameters: - out_suffix (str) – Suffix to append to original filename
- outdir (str) – Path to output directory
- force_rerun (bool) – If structure should be re-cleaned if a clean file exists already
- remove_atom_alt (bool) – Remove alternate positions
- keep_atom_alt_id (str) – If removing alternate positions, which alternate ID to keep
- remove_atom_hydrogen (bool) – Remove hydrogen atoms
- add_atom_occ (bool) – Add atom occupancy fields if not present
- remove_res_hetero (bool) – Remove all HETATMs
- keep_chemicals (str, list) – If removing HETATMs, keep specified chemical names
- keep_res_only (str, list) – Keep ONLY specified resnames, deletes everything else!
- add_chain_id_if_empty (str) – Add a chain ID if not present
- keep_chains (str, list) – Keep only these chains
Returns: Path to cleaned PDB file
Return type: str
-
file_type= None¶ str – Type of structure file
-
find_disulfide_bridges(threshold=3.0)[source]¶ Run Biopython’s search_ss_bonds to find potential disulfide bridges for each chain and store in ChainProp.
-
get_dict_with_chain(chain, only_keys=None, chain_keys=None, exclude_attributes=None, df_format=False)[source]¶ get_dict method which incorporates attributes found in a specific chain. Does not overwrite any attributes in the original StructProp.
Parameters: - chain –
- only_keys –
- chain_keys –
- exclude_attributes –
- df_format –
Returns: attributes of StructProp + the chain specified
Return type: dict
-
get_dssp_annotations(outdir, force_rerun=False)[source]¶ Run DSSP on this structure and store the DSSP annotations in the corresponding ChainProp SeqRecords
Calculations are stored in the ChainProp’s
letter_annotationsat the following keys:SS-dsspRSA-dsspASA-dsspPHI-dsspPSI-dssp
Parameters: - outdir (str) – Path to where DSSP dataframe will be stored.
- force_rerun (bool) – If DSSP results should be recalculated
Todo
- Also parse global properties, like total accessible surface area. Don’t think Biopython parses those?
-
get_freesasa_annotations(outdir, include_hetatms=False, force_rerun=False)[source]¶ Run
freesasaon this structure and store the calculated properties in the corresponding ChainProps
-
get_msms_annotations(outdir, force_rerun=False)[source]¶ Run MSMS on this structure and store the residue depths/ca depths in the corresponding ChainProp SeqRecords
-
get_structure_seqs(model)[source]¶ Gather chain sequences and store in their corresponding
ChainPropobjects in thechainsattribute.Parameters: model (Model) – Biopython Model object of the structure you would like to parse
-
is_experimental= None¶ bool – Flag to note if this structure is an experimental model or a homology model
-
load_structure_path(structure_path, file_type)[source]¶ Load a structure file and provide pointers to its location
Parameters: - structure_path (str) – Path to structure file
- file_type (str) – Type of structure file
-
mapped_chains= None¶ list – A simple list of chain IDs (strings) that will be used to subset analyses
-
parse_structure(store_in_memory=False)[source]¶ Read the 3D coordinates of a structure file and return it as a Biopython Structure object. Also create ChainProp objects in the chains attribute for each chain in the first model.
Parameters: store_in_memory (bool) – If the Biopython Structure object should be stored in the attribute structure.Returns: Biopython Structure object Return type: Structure
-
parsed= None¶ bool – Simple flag to track if this structure has had its structure + chain sequences parsed
-
structure= None¶ Structure – Biopython Structure object, only used if
store_in_memoryoption ofparse_structureis set to True
-
structure_file= None¶ str – Name of the structure file
-
view_structure(only_chains=None, opacity=1.0, recolor=False, gui=False)[source]¶ Use NGLviewer to display a structure in a Jupyter notebook
Parameters: - only_chains (str, list) – Chain ID or IDs to display
- opacity (float) – Opacity of the structure
- recolor (bool) – If structure should be cleaned and recolored to silver
- gui (bool) – If the NGLview GUI should show up
Returns: NGLviewer object
- Provide access to the 3D coordinates using a Biopython Structure object through the method